Precision Modification of Bio-molecules
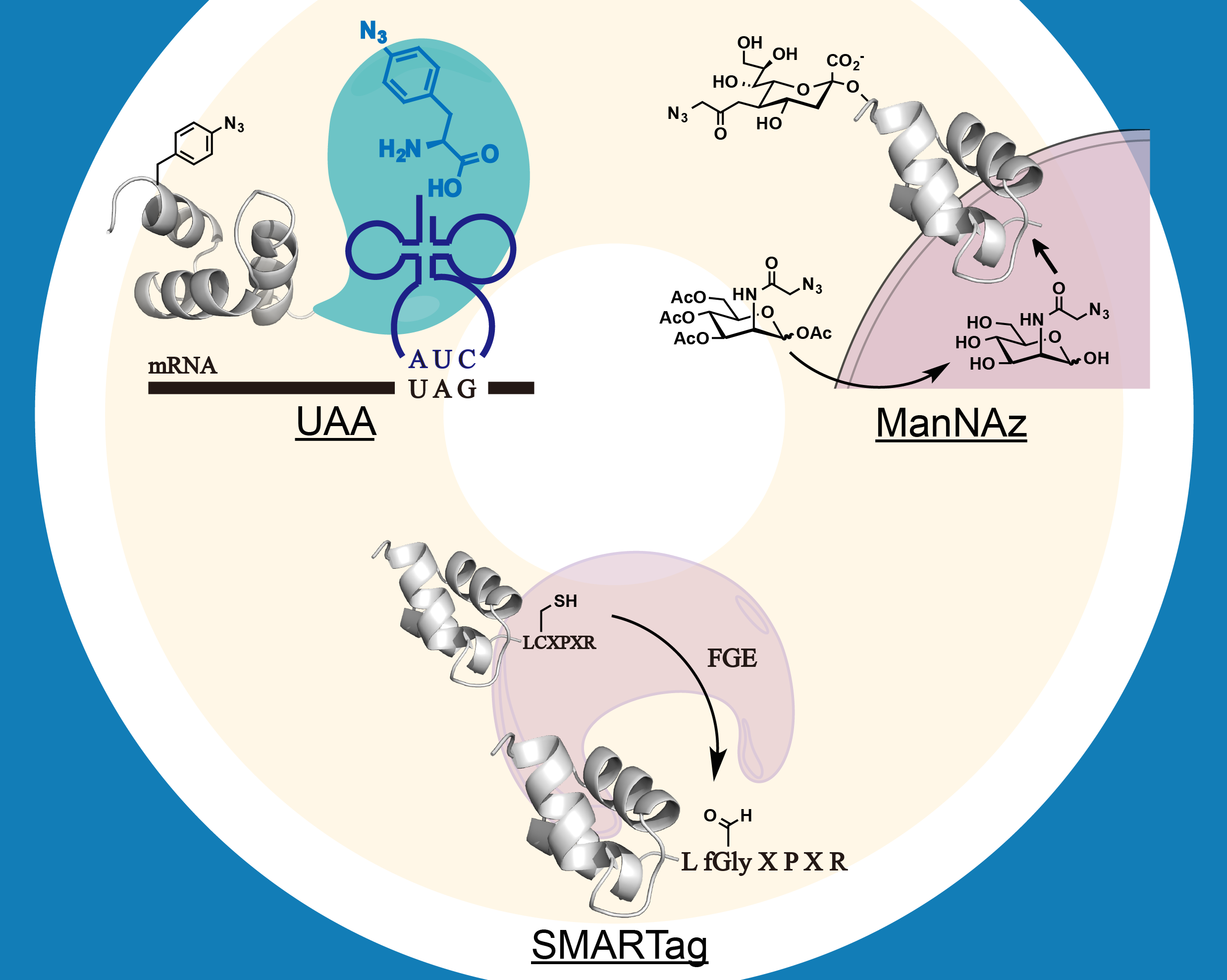
The ability to site-specifically introduce distinct chemical moieties (e.g. unnatural amino acid, unnatural saccharide derivatives) into biomolecules affords a powerful tool to investigate and manipulate biological systems. Here, we will expand the tool set for this purpose. We are interested in (1) adding new building blocks with novel chemical, biological, and physical properties in different biological systems; (2) enhancing the performance of chemical biological tools in various species; (3) using these tools to better understand and ultimately control various biological processes; and (4) exploring the therapeutic utilities of these tools in the context of cancer, autoimmune, and metabolic diseases.
First-in-Class Therapies for Bone Diseases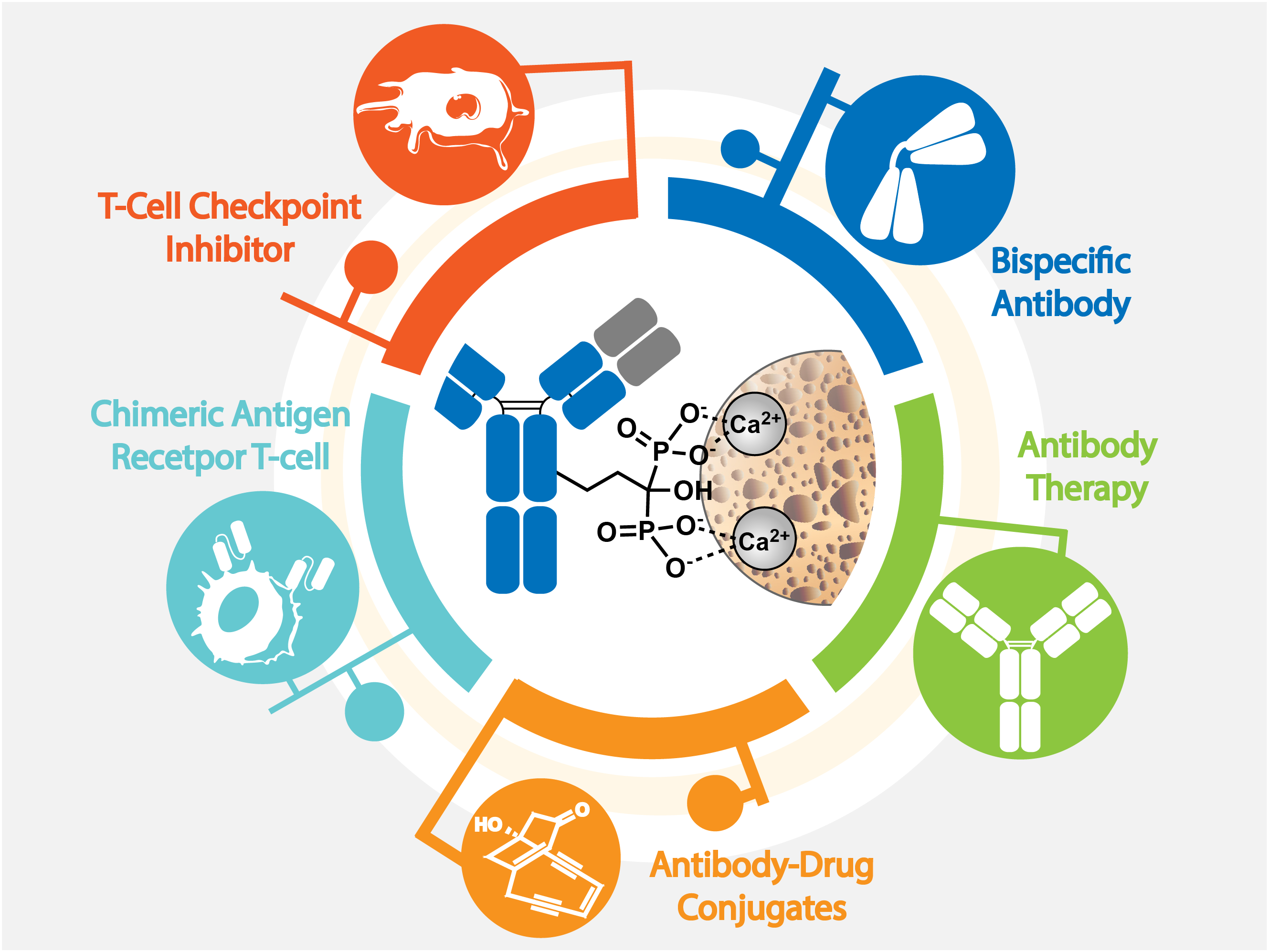
The skeletal system provides structural support and maintains mineral homeostasis, but it also plays a critical role in the progression of diseases such as cancer metastasis, autoimmune disorders, and hematologic malignancies. Breast and prostate cancers are the most common sources of bone metastases; over 47% of initial metastatic events occur in bone, and about 80% of prostate cancer metastases are confined to the skeleton. However, effective drug delivery to bone remains a challenge, limiting current treatment efficacy. To address this, my lab developed BonTarg, a bone-targeting platform that enables site-specific conjugation of bone-homing moieties to therapeutic antibodies. This approach enhances antibody accumulation in bone metastases while minimizing off-target effects. We also identified a novel glyco-immune checkpoint in the bone tumor microenvironment and showed that antibody blockade suppresses tumor-induced bone damage and boosts T cell responses. These advances support a dual strategy—targeted delivery and immune modulation—for treating bone-metastatic cancers. We are further expanding this platform to deliver small molecules, epigenetic inhibitors, bispecific antibodies, and cell therapies to the bone niche.
Machine Learning-Based Protein Evolution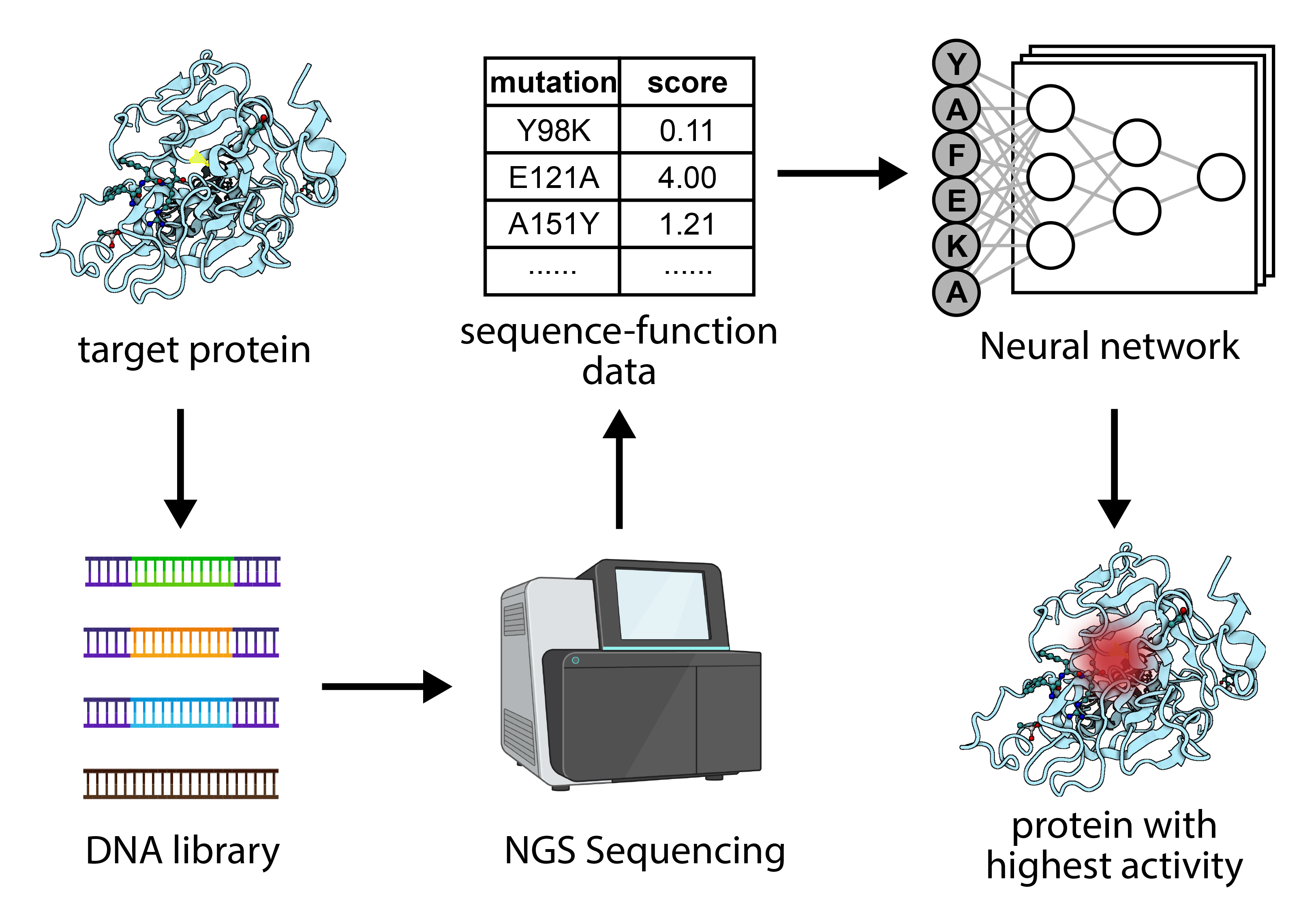
Machine learning has brought about a paradigm shift in protein evolution, facilitating swift and precise projections of protein structures, functions, and interactions. This advancement accelerates the innovation of novel enzymes, therapeutic agents, and biomolecules endowed with amplified attributes and custom functionalities. A present challenge lies in swiftly amassing sequence-function data for a target protein. To address this, we are devising a comprehensive approach that employs next-generation sequencing (NGS) to acquire functional information for an array of target proteins. Integrating this resultant sequence-function dataset with neural networks, we aim to engineer new proteins possessing attributes surpassing those found in natural sequences.
Biomedical Probes for Noninvasive Deep-Tissue Imaging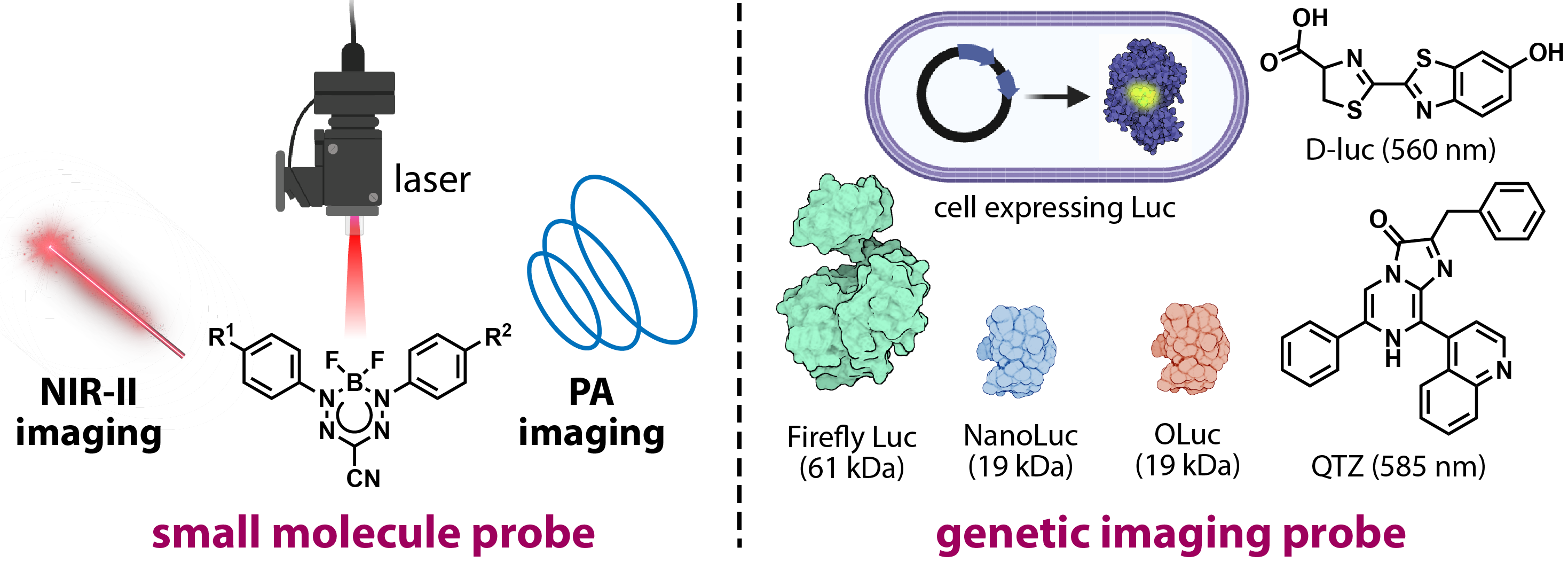
Compared to traditional imaging, NIR-II, bioluminescence, and photoacoustic imaging offer deeper tissue penetration with reduced photon scattering and autofluorescence, making them powerful tools for noninvasive biomedical imaging. However, probes suitable for these modalities remain limited, especially those capable of crossing the blood-brain barrier or entering the lymphatic system. To address this gap, we are developing advanced functional imaging probes for in vivo NIR-II imaging and photoacoustic computed tomography. These probes are designed to achieve high spatial resolution, target specificity, and favorable biodistribution, enabling precise and dynamic visualization of deep tissues in both research and potential clinical applications.
Acknowledgement of Support
Many thanks to the following funding sources for making our work possible.








